Modelling Host-Microbiome Evolution
Who am I
- Background: Mathematical evolution and ecology
- PhD: BCB, UI, Scott Nuismer ~ Coevolutionary Theory
- Postdoc 1: MSU, Gideon Bradburd ~ Host-Parasite Coevolution
- Postdoc 2: UO, Brendan Bohannan ~ Host-Microbiome Evolution
- Current: KiTE postdoctoral fellow with Schulenburg Group at Kiel University
- Host-Microbiome Evolution
- Two Secondments:
- Regional non-academic (industry)
- International academic
- Hosted by Brendan Bohannan at UO
- Purpose: travel to share science and meet scientists
We live in a microbial world
Microbes interact with hosts
metabolism / digestion / nutrition

immune response / pathogen defense
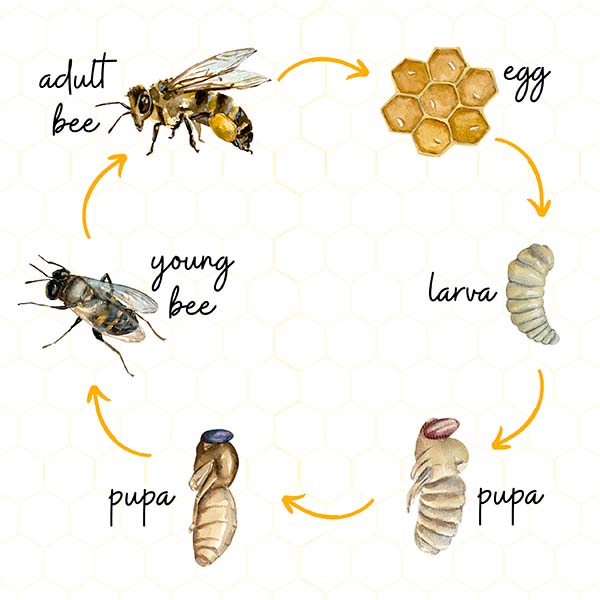
development / life-history / phenology

behavior / neurophysiology
Microbiomes mediate host trait variation
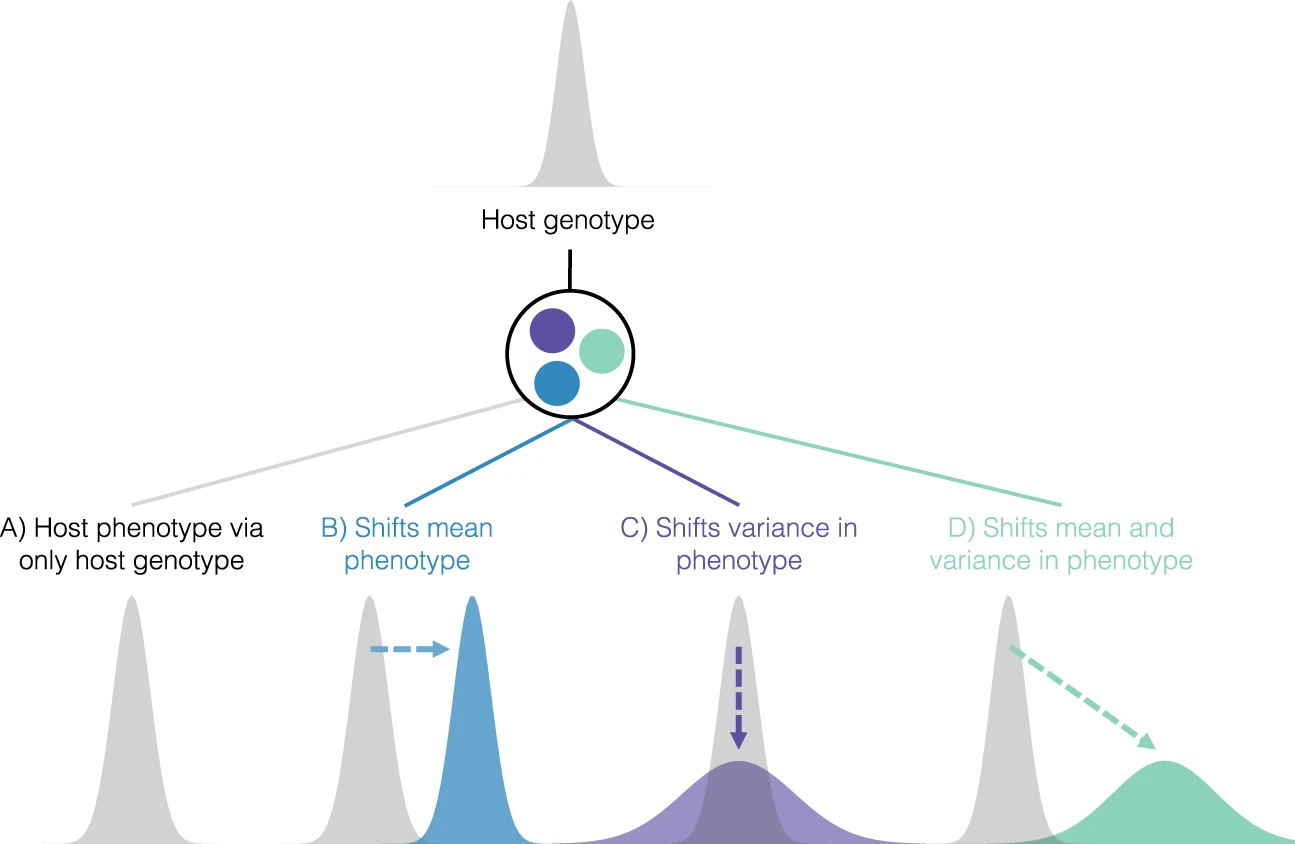
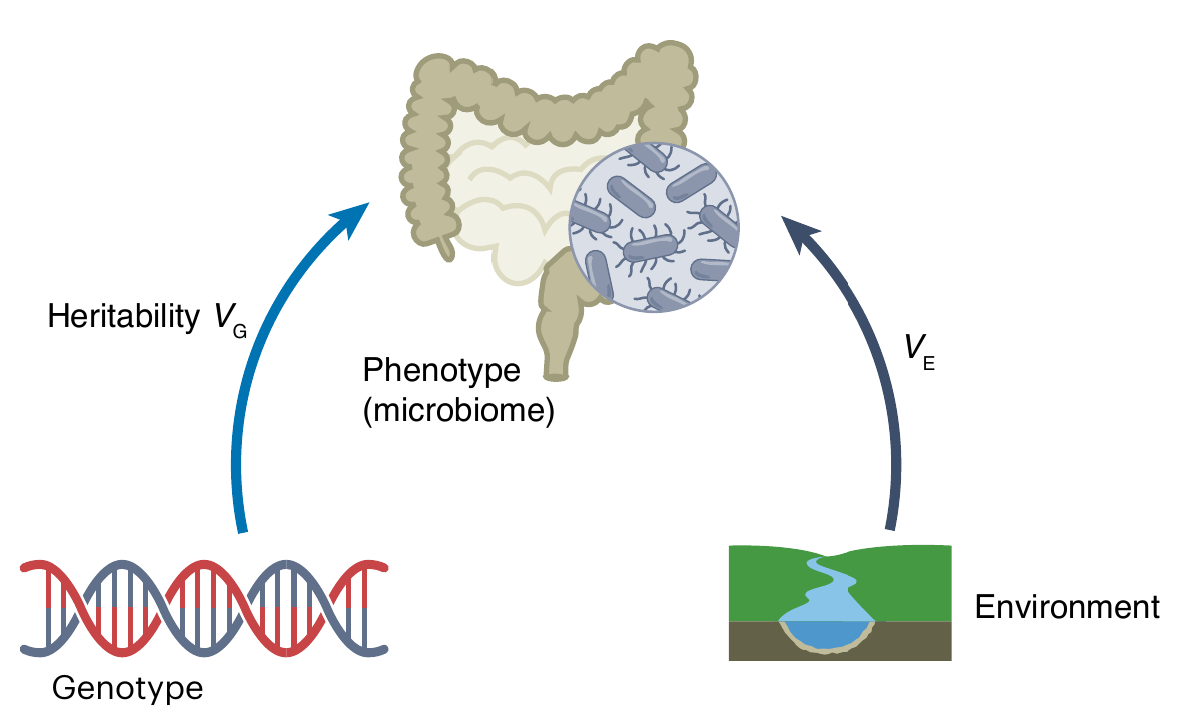
Microbiomes are heritable
- Fraction of attributes heritable: ~5–40%
- relative abundance, alpha diversity, etc …
- Broad sense heritability: H² ≈ 0.01 - 0.5
- perfect heritability = 1.0
Microbiome inheritance modes
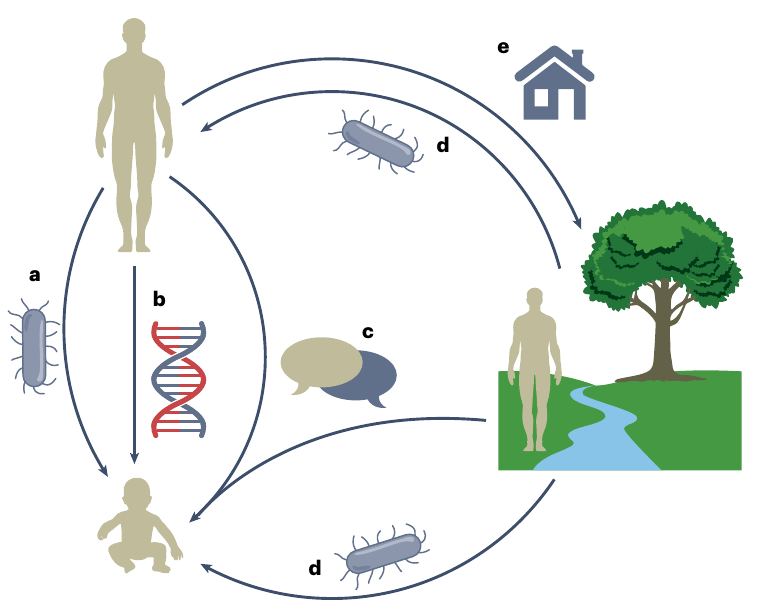
Morris & Bohannan (2024), Nat. Micr.
- Direct parent-offspring transmission
- Host genes associated with microbes
- Social transmission
- Shared environment
- Habitat modification
Can Microbiomes facilitate adaptation?
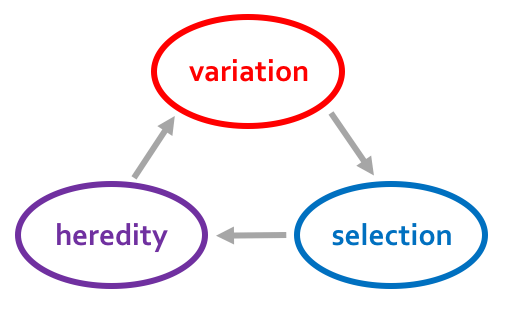
But Microbial Inheritance not Mendelian …
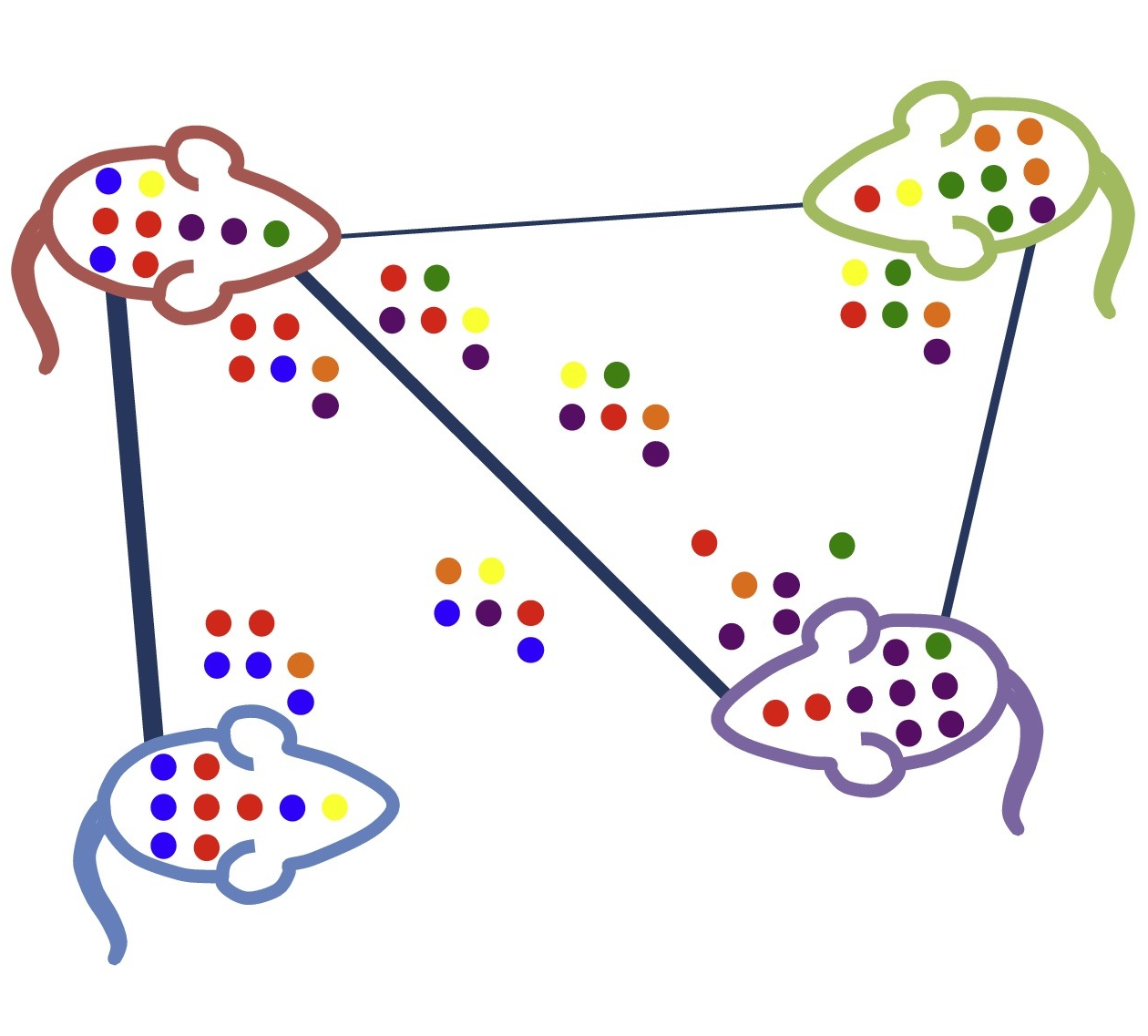
Community assembly (microbial dispersal, within-host selection, etc …)
Problem: Complicated!
Need: Simple general frameworks
Q: Can we apply existing frameworks?
Extended inheritance frameworks
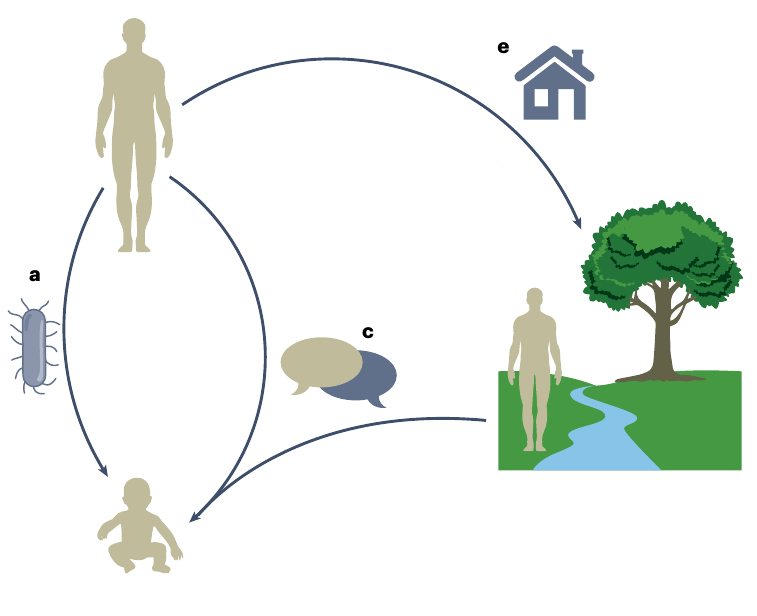
Morris & Bohannan (2024), Nat. Micr.
- Maternal effect
- Indirect genetic effect
- Niche construction
Maternal Effect
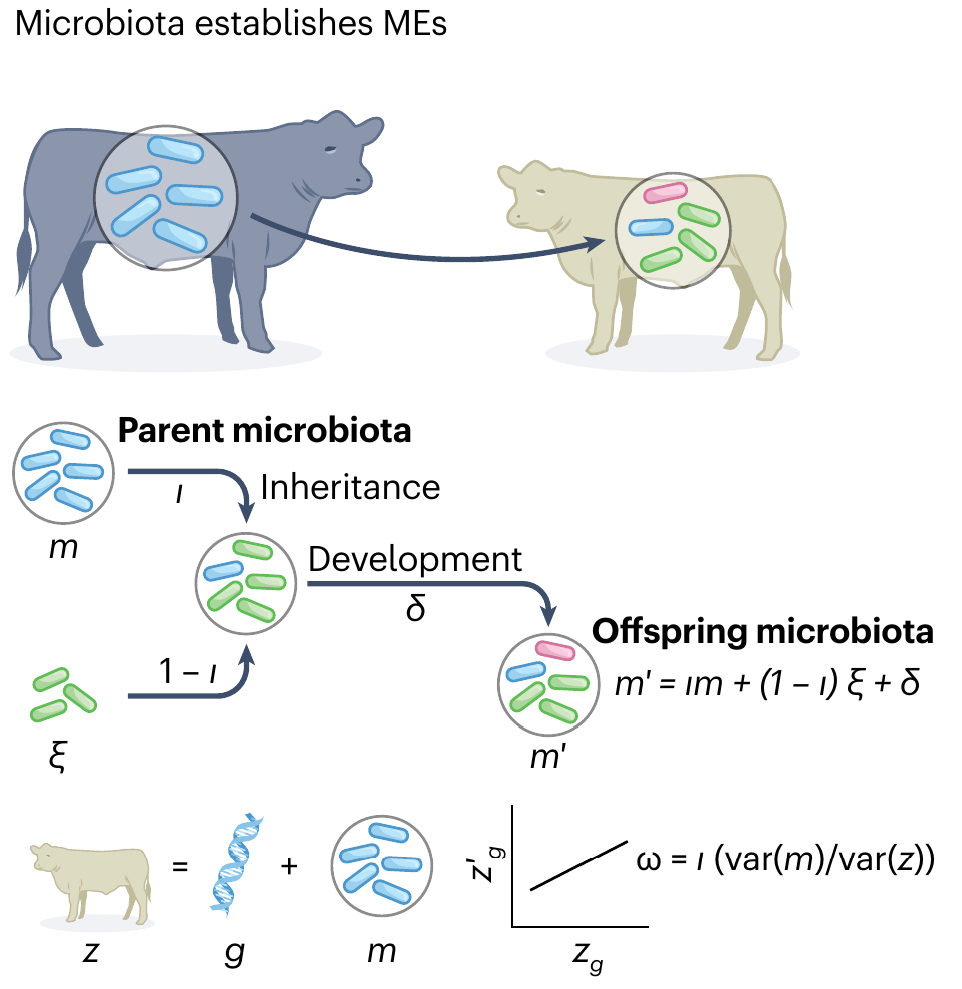
Week et al. (2024), Nat. Evo. Eco.
Definition: Parent trait explains offspring trait
(controlling for genetic variation)
Indirect Genetic Effect
Definition: Genotype of one individual influences trait of another individual
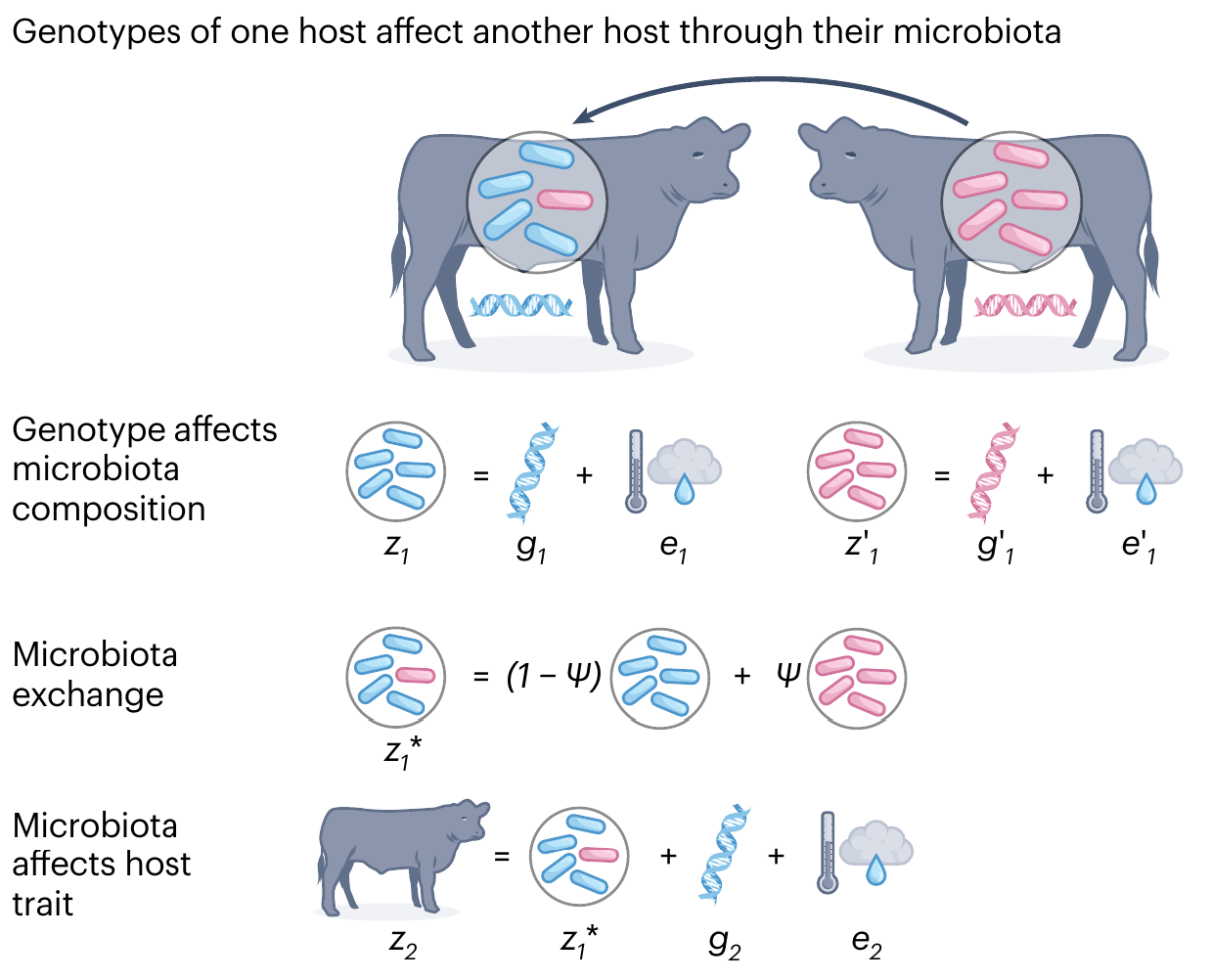
Week et al. (2024), Nat. Evo. Eco.
Niche Construction
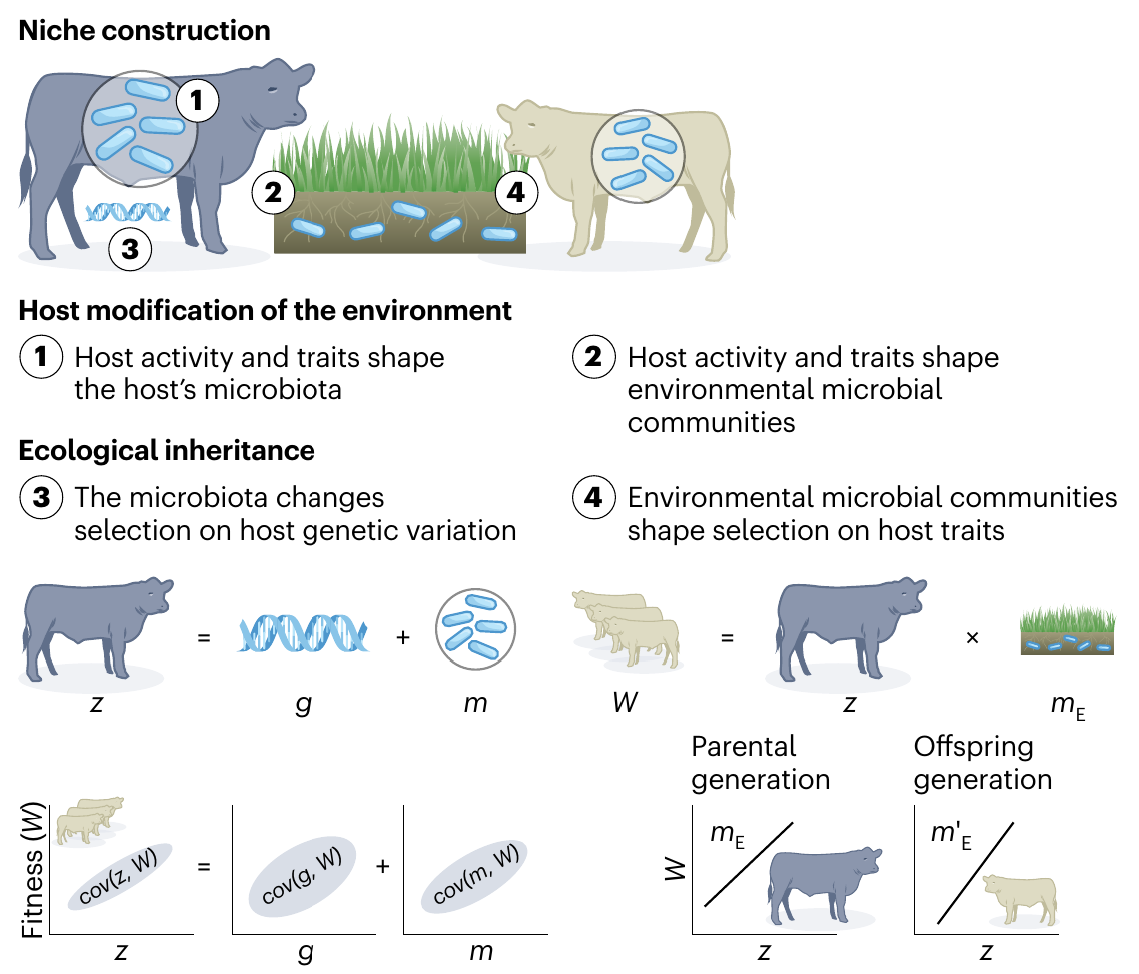
Week et al. (2024), Nat. Evo. Eco.
Definition 1:
Organism activity alters environment
Definition 2:
Organism activity alters selection pressures
Two perspectives:
Microbiome as environment
Microbiome modifying environment
Conclusion
PROS
- Maternal Effects ~ vertical parent-offspring inheritance
- Indirect Genetic Effects ~ horizontal social inheritance
- Niche Construction ~ interaction of microbiomes and selection
CONS
- Each only cover one aspect
- Microbiome inheritance follows several modes
- Need: unified/sythetic framework
Unified Framework for Extended Inheritance
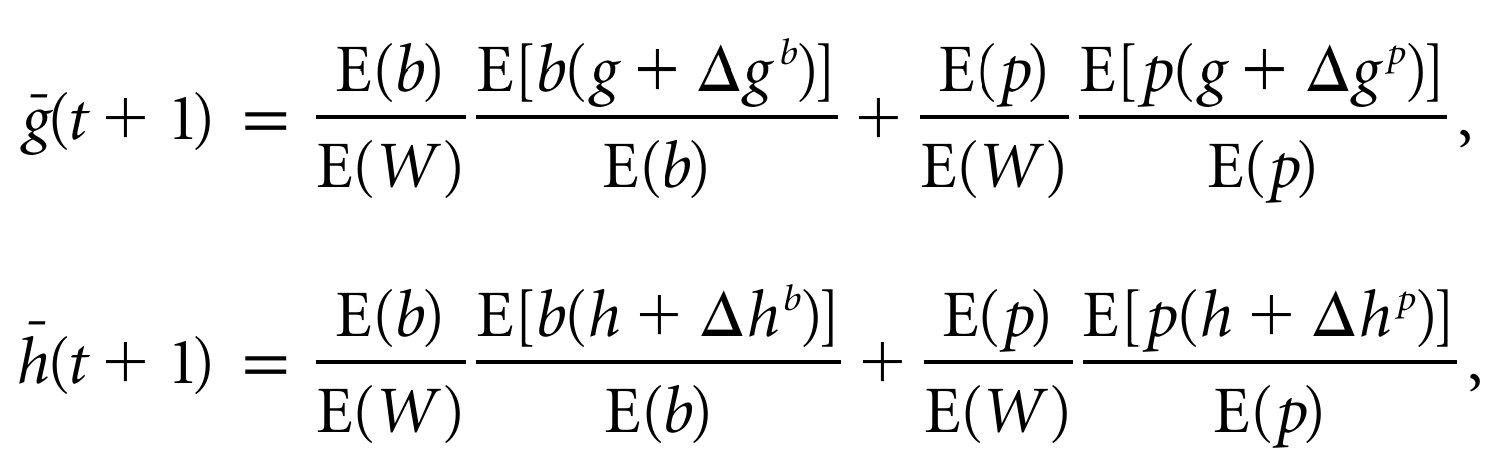
- Models: ME’s, IGE’s, etc.
- Limitation: requires assumptions on
- transmission
- fidelity
- mutability
Framework for Microbiome Inheritance
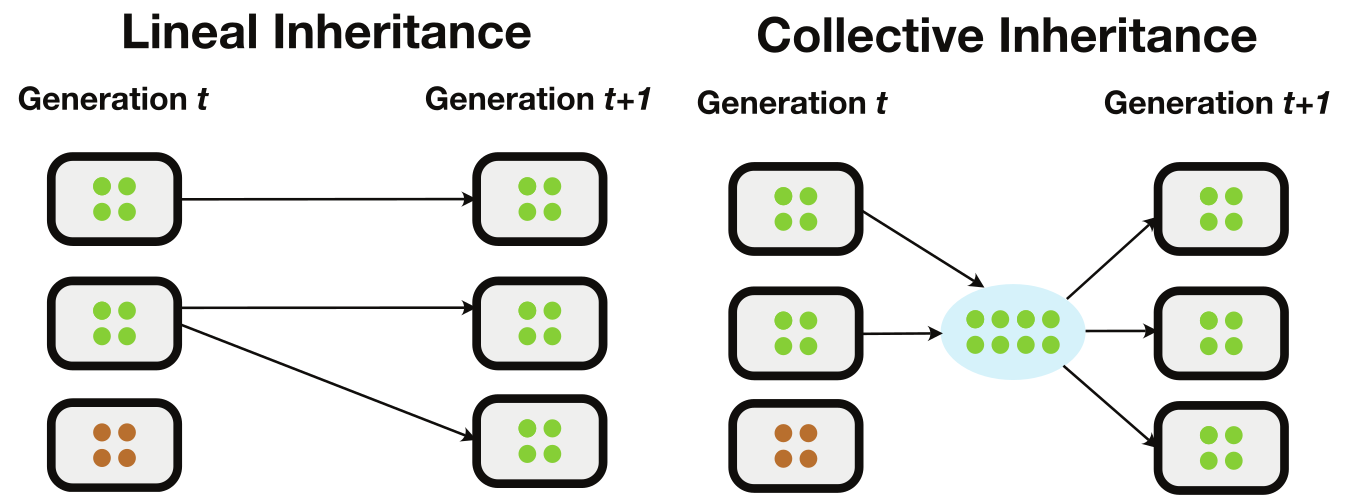
- Collective inheritance facilitates adaptation
- Advantage: Potential to explain adaptation without transmission details
A Gap
Done: Collective inheritance applied to gene-centric perspective 🦠 🚚 🧬
Gap: Connect concepts of microbiome inheritance and host trait variance to predict microbiome-mediated adaptation
microbiome inheritance
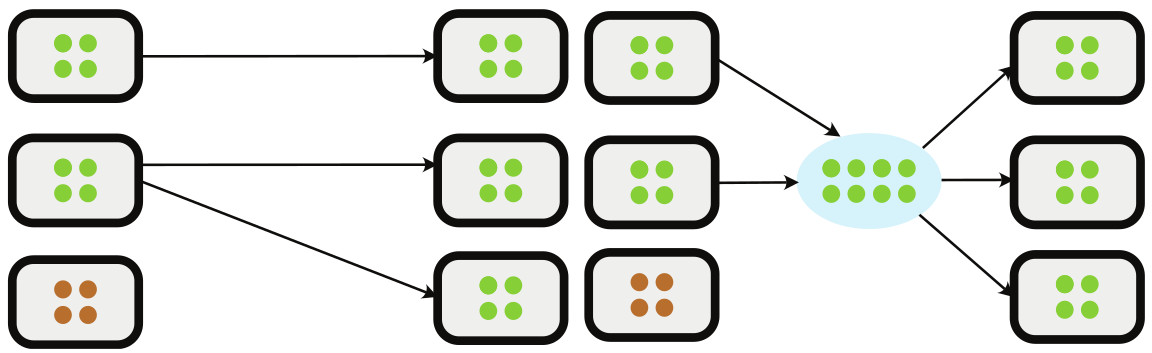
host trait adaptation
Is a Quantitative Genetic Approach Useful?
Explains genetic adaptation of complex traits ✅
Can account for trait variation explained by microbiomes ✅
Adaptation predictions assume Mendelian inheritance ❌
- Interface with microbiome inheritance to close gap ✅
Classical Partitioning of Trait Variation
mini review of quantitative genetics
- \(P = G + E\)
- \(G = G_A + R\)
- Adaptation Prediction:
- Resp. to Sel. = \(G_A\) × Sel. Grad.
Additive Genetic Variance \(G_A\)
Partitioning of Variation including Microbiome
- \(P_A = G_A + M_A + C_A\)
- \(M_A=\) microbial variation
- \(C_A=\) gene-microbe covariance
- Adaptation Prediction:
- Resp. to Sel. = \(P_A\) × Sel. Grad. ?
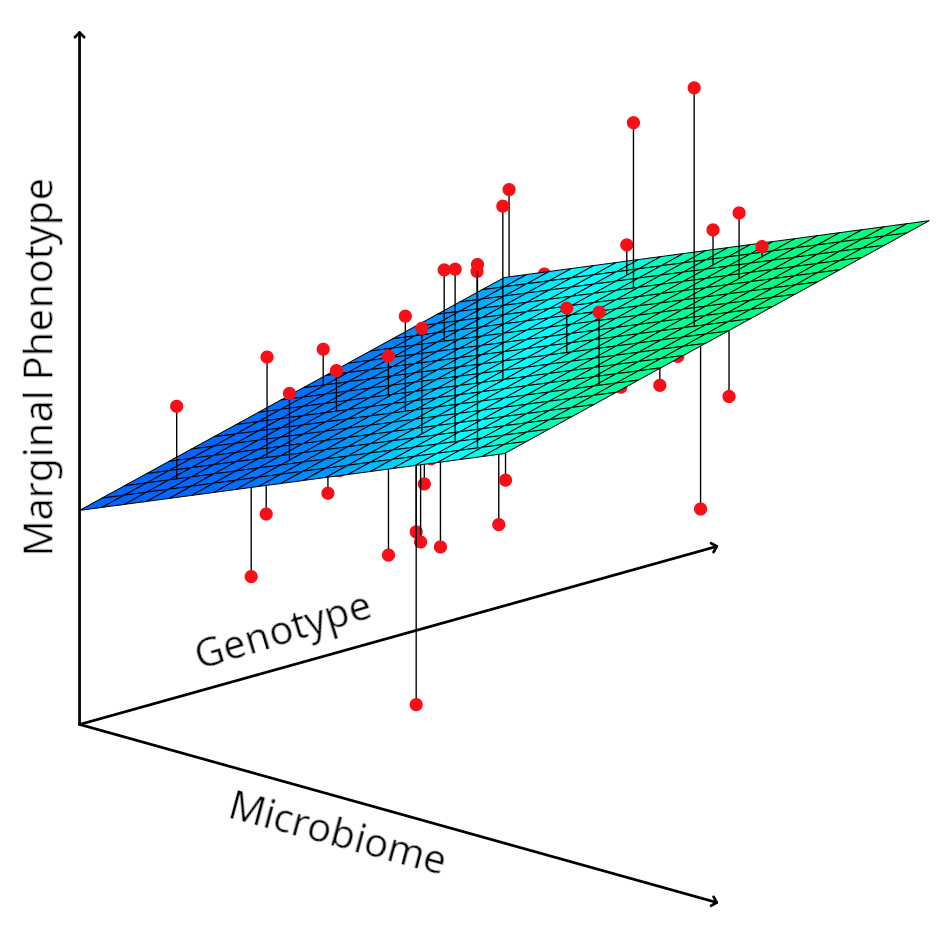
Not all microbes contribute to adaptation
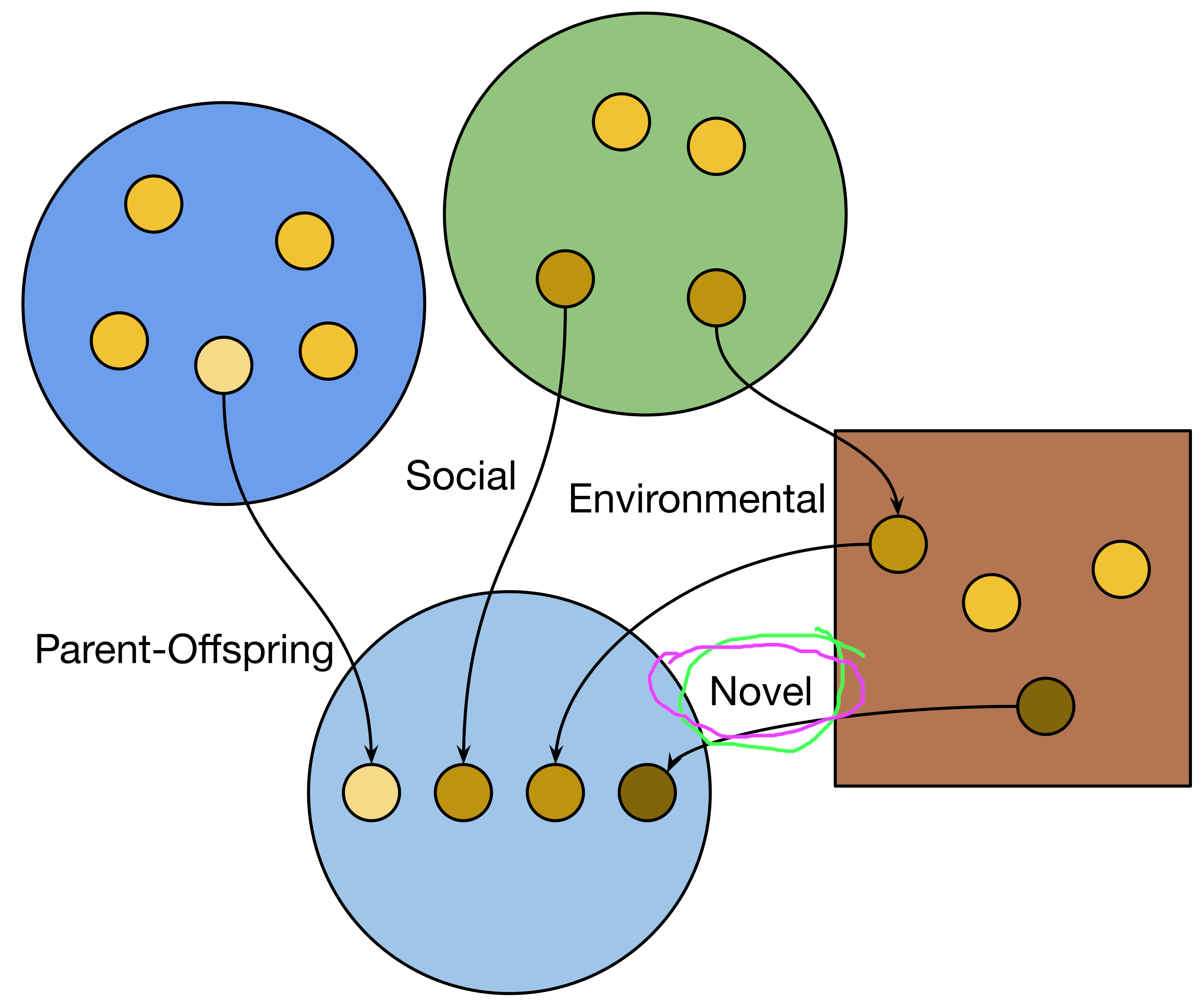
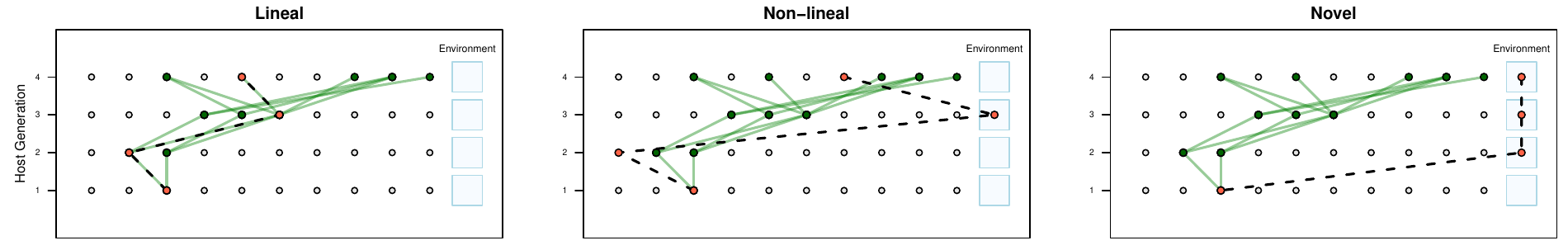
Ancestral Concordance Classification
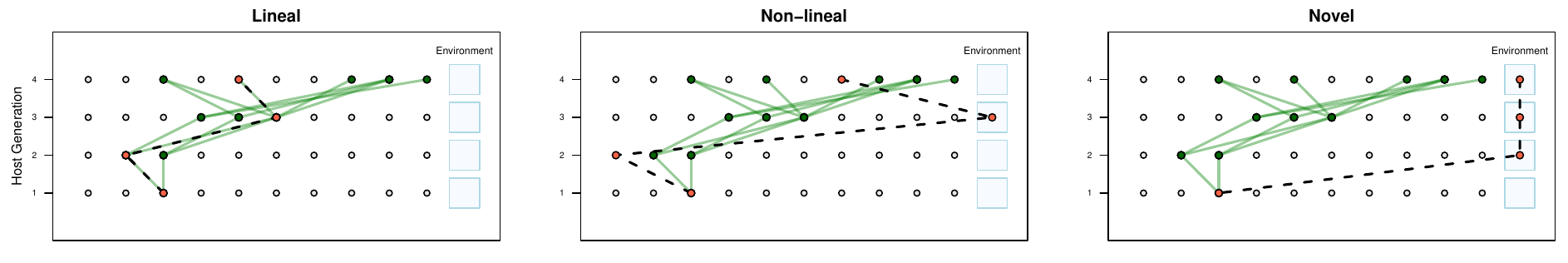
Lineal Microbes:
Ancestry overlaps with host ancestry
Non-lineal Microbes:
Ancestry in host population
Novel Microbes:
No previous ancestry with host population
classifies individual microbial cells
Partitioning Var. by Ancestral Concordance

\(M_L=\) lineal variance
\(M_N=\) non-lineal variance
\(M_V=\) noVel variance
- (Additive Phenotypic Variation) \(P_A = G_A + {\color{magenta}{M_A}} + C_A\)
- (Additive Microbial Variation) \({\color{magenta}{M_A}} = M_L + M_N + M_V + {\color{gray}{C_{LNV}} }\)
Hypothesis
- Adaptation:
- Lineal: \(M_L\) most likely
- Non-Lineal: \(M_N\) less likely
- NoVel: \(M_V\) no contribution
- How to test:
- Host Pedigree
- Metagenomic Data for Microbiomes of Hosts in Pedigree
- Strain Resolved
- Classify Microbes by Ancestral Concordance
- Partition host trait variance
- Selection Experiment
- Predict response including different combinations of \(M_L,M_N,M_V\)
- Measure response to selection and assess predictions
Simulations
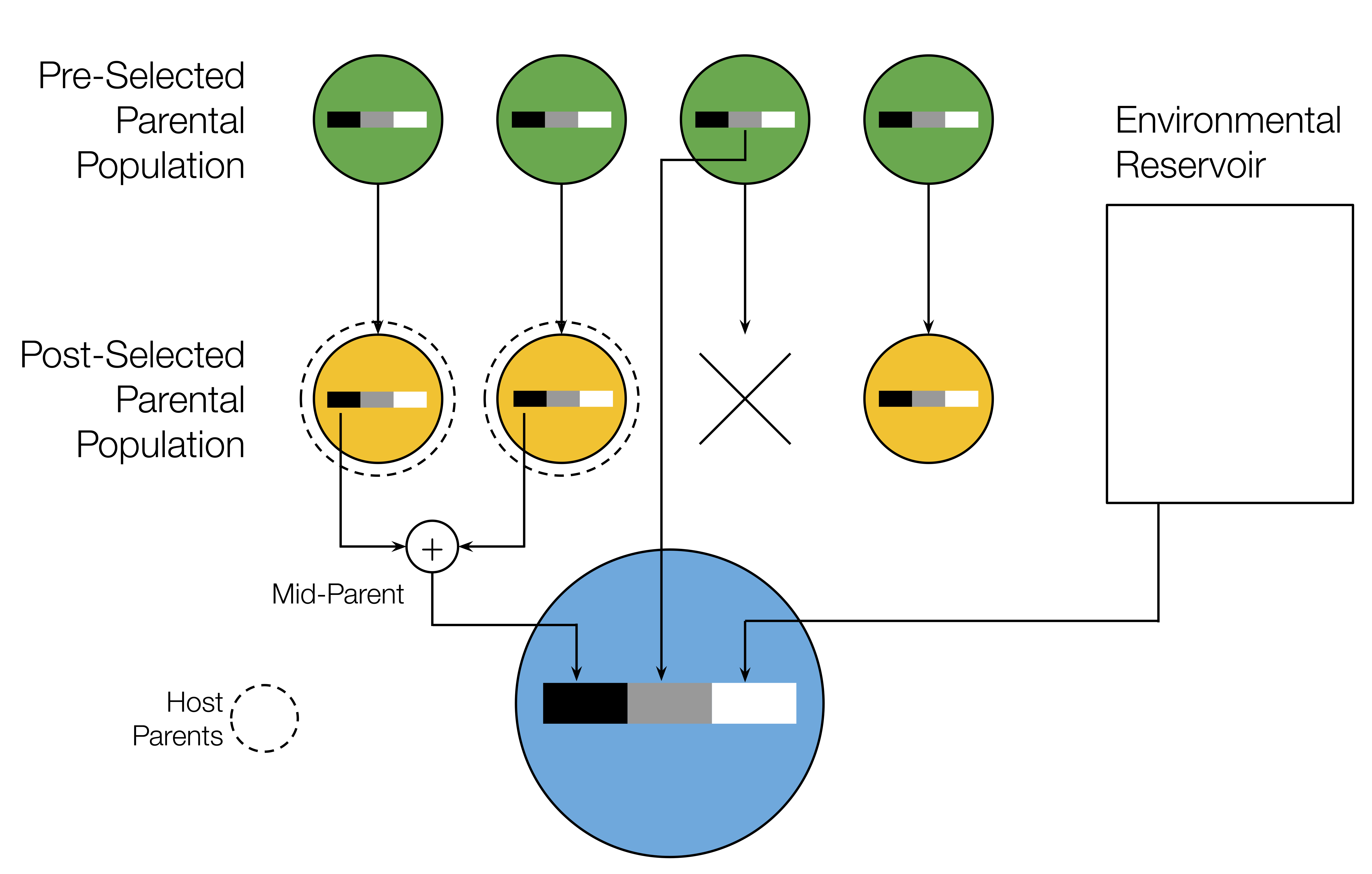
[ 🖤 lineal | 🩶 non-lineal | 🤍 novel ]
Simple inheritance model:
- Lineal abundances
- sampled from parents
- Non-lineal abundances
- sampled from random host
- before or after selection
- Novel abundances
- drawn independently
Additive host trait:
- \(z=\sum_i\alpha_i\,n_i\)
Simulations
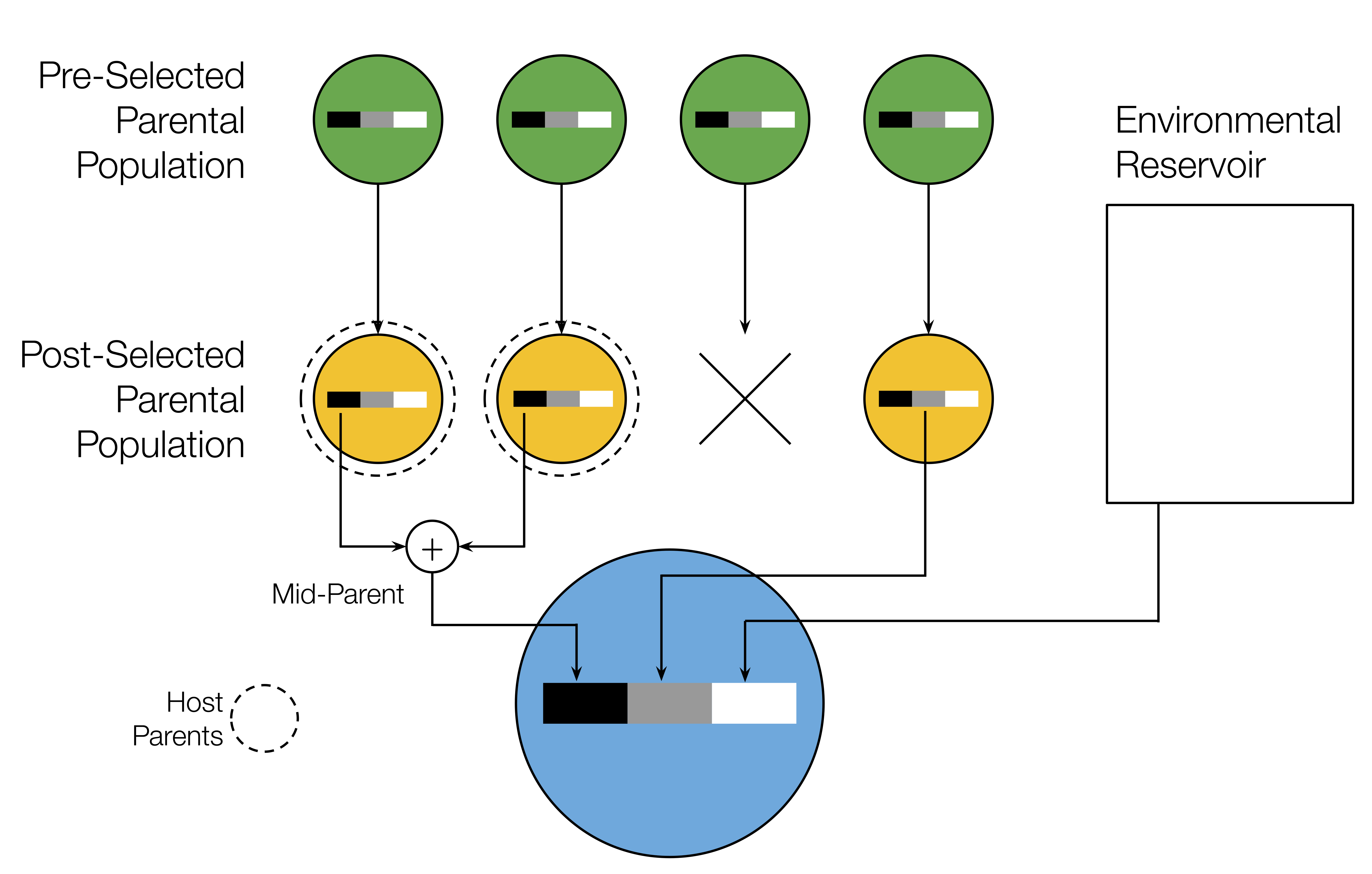
[ 🖤 lineal | 🩶 non-lineal | 🤍 novel ]
Simple inheritance model:
- Lineal abundances
- sampled from parents
- Non-lineal abundances
- sampled from random host
- before or after selection
- Novel abundances
- drawn independently
Additive host trait:
- \(z=\sum_i\alpha_i\,n_i\)
Results
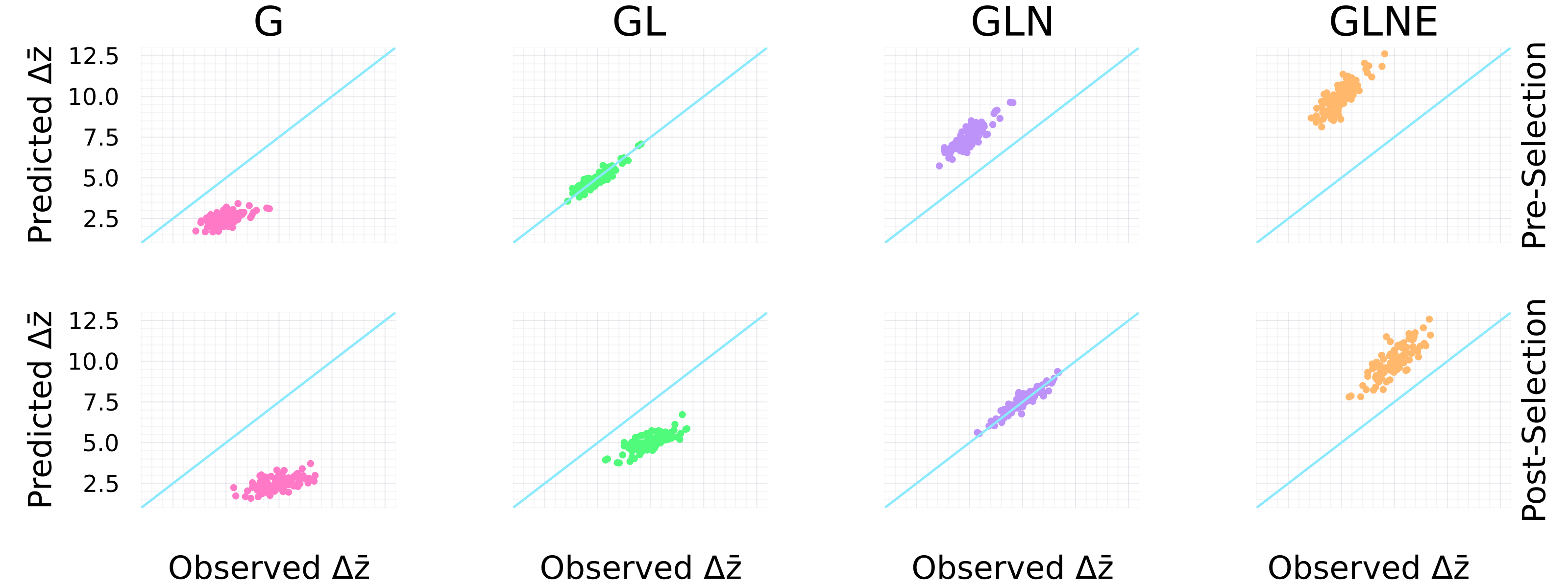
\(\pmb\Delta\mathbf{\bar z}\) = Resp. to Selection, G = Gen. var, L = Lineal, N = Non-lineal, V = NoVel
Conclusion: include only microbe variation shaped by selection
The Selective Microbes and Heritability
Definition: Subset of microbes that contribute to host adaptation
- \(M_A^\psi\) = trait variance explained by variance of selectives
- \(C_A^\psi\) = trait variance explained by covariance of selectives and genes
By definition: (Resp. to Sel.) = (\(G_A+M_A^\psi+C_A^\psi\)) × (Sel. Grad.)
Without microbe fx: (Resp. to Sel.) = (\(G_A\)) × (Sel. Grad.)
Narrow-sense heritability: \(h^2\) \(=\)\(\frac{G_A}{P}\), Resp. to Sel.: \(R=\) \(h^2\) \(S\)
Extending Heritability
Narrow-sense heritability: \[h^2=\frac{G_A}{P}\]
\(R=h^2\,S\)
(w/o micr. fx)
Narrow-sense total transmissibility:
\[t^2=\frac{M_A^\psi+C_A^\psi}{P}\]
\(R=t^2\,S\)
(w micr. fx)
Read all about it!
Thanks Co-Authors




What about Theory?
- Quant. Gen. approach:
- Statistical framework
- Aimed at empirical studies
- Still very complicated
- Initial theoretical insights:
- Simple models
- Motivates some current projects…
Microbe-Mediated Host Rescue

MPI Plön
Scenario: Doomed host population
Microbiome: Presence/absence single microbe
- Probability of lineal inheritance
- Rate of social transmission
Fitness: Host gene determines microbe benefit
Results: Delayed host extinction time analysis
Birth-Death Diffusion \(dn=[(r+m)\,n+\psi]\,dt+\sqrt{n}\,dB\)
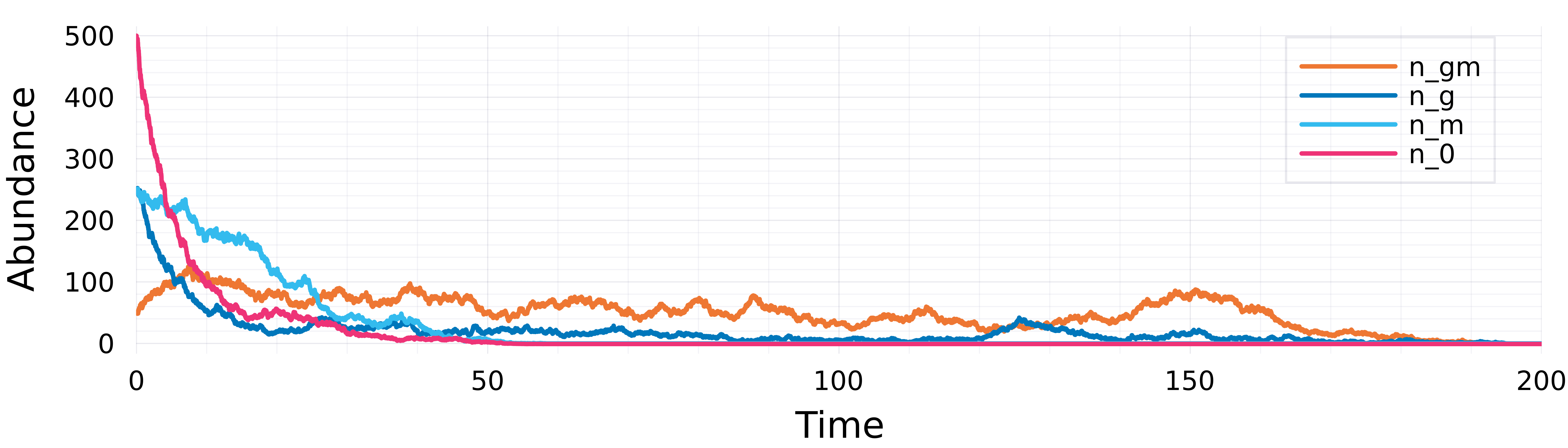
Microbiome-Mediated Niche Construction

Kiel University
- Mesocosm experiments:
- C. elegans shape microbiome in compost
- Compost conditions degrade
- Microbiome manipulation buffers against \(\Delta E\)
- Quant. Gen. Theory
- Modify \(\Delta\bar z=G\beta\)
- Modelling framework (instead of statistical)
- Microbiome community dynamics ~ mutation
- Feedback with external microbiome
- External microbiome changes effect of env.
- Modify \(\Delta\bar z=G\beta\)
Q: What eco-evo processes and what host-microbiome attributes shape adaptation via niche construction?
Multilevel Selection Framework

MPI Plön
- Mesocosm experiments:
- C. elegans shape microbiome in compost
- Compost conditions degrade
- Microbiome manipulation buffers against \(\Delta E\)
- MLS Model
- Modify \(\Delta\bar z=G\beta\)
- Modelling framework (instead of statistical)
- Microbiome transmission + dynamics ~ mutation
- Feedback with external microbiome
- External microbiome changes effect of env.
- Modify \(\Delta\bar z=G\beta\)
Q: What eco-evo processes and what host-microbiome attributes shape adaptation via niche construction?
Entangled Banks

Social Microbiome Metacommunity Framework
University of Oxford
clusters
bridges